GWAS simulation setting - constant value vs random variable
update at 19/Jan/2024
I revisit this post after reading Section 4.4 in the book Casella and Berger, Statistical Inference (2001)
===================
Simulation is a common method to study (bio)statistics, like quantitavie genetics or gwas (genome-wide association study).
GWAS simulation setting
A common way to simulate a phenotype based on one simulated SNP (e.g. Wu et al. Genome Biology 2017) is:
y = g + e; g = wu
- y is a simulated phenotype
- w is a simulated standardized (mean 0 and variance 1) genotype, calculated by w = (x-2f)/sqrt(2f(1-f))
- x is a simulated genotype, generated from Bin(2, f)
- f is the minor allele frequency, generated from Uni(0.1, 0.5)
- u is the effect size per standardized genotype, generated from N(0,1)
- g is the genetic value, calculated by wu
- e is the error term, generated from N(0, var(g)(1/h2 - 1))
- h2 is the variance explained by the genotype (heritability)
Constant value vs random variable
In this setting,
- y, w, x, and e are random variables
- h2 is a parameter, which is usually set manually
Importantly note that
f and u are also constant values, although it is generated randomly
var(g) = var(wu) = u^2var(w) = u^2
E(var(gu)) = E(u^2) = var(u) + E(u)^2 = 1 + 0^2 = 1
R code
We can verify the above in R
set.seed(7)
n=10000; rep=500; res=c()
for(repindex in 1:rep){
f = runif(1, 0.1, 0.5)
x = rbinom(n, 2, f)
w = (x-2*f)/sqrt(2*f*(1-f))
u = rnorm(1)
g = w*u
res = c(res, var(g))
}
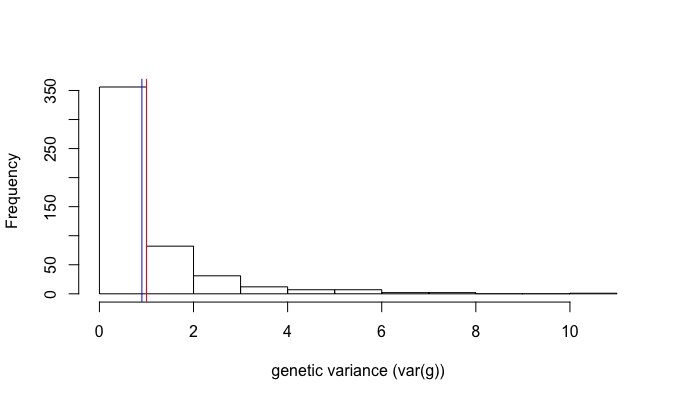
hist(res, main = "", xlab="genetic variance (var(g))") ### emprical distribution
abline(v=1, col="red") ### expected value
abline(v=mean(res), col="blue") ### emprical value